Current Projects
Optimizing eradication of Brown Treesnakes on Guam
Fall 2022 - Present, USGS WA Cooperative Fish & Wildlife Unit
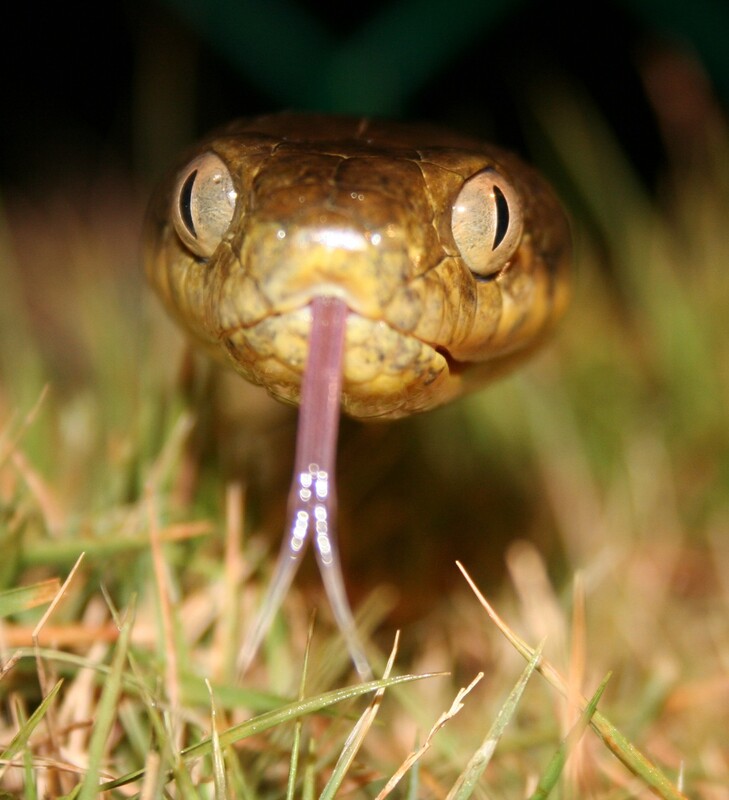
Brown Treesnakes are an invasive species on the island of Guam, unintentionally introduced in the late 1940s by US military transports coming from its native range in New Guinea. Guam does not have any native snake species, and as the Brown Treesnake population grew over the last 80 years, many of its native bird species were driven to local extinction due to the increased predation. In the 1980s, Dr. Julie Savidge determined that the bird declines were primarily being caused by Brown Treesnakes, which was the first time that an invasive snake had been known to cause ecosystem-wide disturbances like local extinctions (Savidge, 1987). In order to protect Guam’s native ecosystem, and because of their other impacts on the human population of Guam (such as causing power outages), scientists and wildlife managers have been working for the last 40 years on improving our knowledge of the Brown Treesnake and creating methods for controlling and eventually eradicating Brown Treesnakes from Guam.
I am contributing to this effort through the USGS WA Cooperative Fish & Wildlife Unit by conducting a simulation experiment to test out different eradication method scenarios on various snake population conditions. The goal of this study is to determine if there is a scenario (or scenarios) that are optimal for a wide range of different population conditions or if the optimal scenario is highly dependent on what is happening with the snake population. If one or a few eradication method scenarios prove to be optimal across a range of population conditions, then I will be able to provide a relatively straightforward recommendation for the eradication method scenario that is likely to be optimal without needing to know more about the underlying snake population dynamics. If the snake population dynamics make a big difference in which eradication method scenario will be optimal, I will hopefully be able to provide specific recommendations for what snake population information (reproduction rates, individual snake growth rates, etc) wildlife managers will need in order to determine what the optimal eradication scenario might be for their specific context.
Savidge, J. A. 1987. Extinction of an island forest avifauna by an introduced snake. Ecology 68:660-668.
I am contributing to this effort through the USGS WA Cooperative Fish & Wildlife Unit by conducting a simulation experiment to test out different eradication method scenarios on various snake population conditions. The goal of this study is to determine if there is a scenario (or scenarios) that are optimal for a wide range of different population conditions or if the optimal scenario is highly dependent on what is happening with the snake population. If one or a few eradication method scenarios prove to be optimal across a range of population conditions, then I will be able to provide a relatively straightforward recommendation for the eradication method scenario that is likely to be optimal without needing to know more about the underlying snake population dynamics. If the snake population dynamics make a big difference in which eradication method scenario will be optimal, I will hopefully be able to provide specific recommendations for what snake population information (reproduction rates, individual snake growth rates, etc) wildlife managers will need in order to determine what the optimal eradication scenario might be for their specific context.
Savidge, J. A. 1987. Extinction of an island forest avifauna by an introduced snake. Ecology 68:660-668.
Past Projects
Spatiotemporal Modeling
Summer 2020 - Summer 2022, University of Washington
For my master's thesis work, I collaborated with principal investigator Cecilia O’Leary at NOAA’s Alaska Fisheries Science Center on improving the statistical model used to determine the spatial distribution of groundfish stocks in the Gulf of Alaska.
For a full description of this project, click here.
For a (less detailed) video version of the project description, click here to see a presentation I gave at the UW SAFS Student Symposium in fall 2021.
For a full description of this project, click here.
For a (less detailed) video version of the project description, click here to see a presentation I gave at the UW SAFS Student Symposium in fall 2021.
Birds ‘n Bogs Citizen science
Spring - Fall 2016, University of Alaska Anchorage
This project was a citizen-science based study of the presence and abundance of migratory boreal bird species local to the bogs around Anchorage. In my role as research assistant, I coordinated citizen science volunteers and had the opportunity to do field work myself, in the form of observational presence/absence studies. I also produced an internal report summarizing our findings for that year.
I participated in the fourth year of a projected ten year project that is currently ongoing, so final results have not yet been published.
I participated in the fourth year of a projected ten year project that is currently ongoing, so final results have not yet been published.
Moose diet DNA analysis
Fall 2016 - Spring 2017, University of Alaska Anchorage
This project was a lab-based project focused on determining the best method for identifying plant species using DNA analysis from moose fecal samples as part of a larger long-term study of moose diet. I applied for and was awarded an undergraduate research grant of $2,000 to fund the project.
The goal of this project was to find unique DNA markers that could be used to determine plant species composition from moose fecal samples, a method that has been used successfully for other species but has not been applied to these specific plant species. Both the plant and fecal samples were collected during a moose diet observational study, so the plant species were known to be represented in the fecal samples. I extracted DNA samples from 13 reference plants and multiple randomly selected fecal samples, followed by PCR amplification of the nrITS 2 region, a subset of the ribosomal cassette that has been found to identify 92.7% of plant samples at a species level (Chen, 2010).
Despite multiple attempts using adjusted methods, I was unable to extract a high enough concentration of DNA from any of the plant samples to analyze unique identifiers. This could have been due to the age of the samples (which were not stored with the intention of preserving DNA), or it could mean that the DNA region I chose to focus on does not produce unique DNA markers in these plants. While it was frustrating to be unable to answer my original research question, it was an excellent learning opportunity, and a good demonstration of the stumbling blocks that are common to the practice of science.
While the larger moose diet study is expected to be published at some point, I don’t expect the DNA sequencing portion of the project to be included in the publication.
Chen, S., H. Yao, J. Han, C. Liu, J. Song, L. Shi, Y. Zhu, X. Ma, T. Gao, X. Pang, K. Luo, Y. Li, X. Li, X. Jia, Y. Lin, and C. Leon. 2010. Validation of the ITS2 Region as a Novel DNA Barcode for Identifying Medicinal Plant Species. PLOS ONE 5:e8613.
The goal of this project was to find unique DNA markers that could be used to determine plant species composition from moose fecal samples, a method that has been used successfully for other species but has not been applied to these specific plant species. Both the plant and fecal samples were collected during a moose diet observational study, so the plant species were known to be represented in the fecal samples. I extracted DNA samples from 13 reference plants and multiple randomly selected fecal samples, followed by PCR amplification of the nrITS 2 region, a subset of the ribosomal cassette that has been found to identify 92.7% of plant samples at a species level (Chen, 2010).
Despite multiple attempts using adjusted methods, I was unable to extract a high enough concentration of DNA from any of the plant samples to analyze unique identifiers. This could have been due to the age of the samples (which were not stored with the intention of preserving DNA), or it could mean that the DNA region I chose to focus on does not produce unique DNA markers in these plants. While it was frustrating to be unable to answer my original research question, it was an excellent learning opportunity, and a good demonstration of the stumbling blocks that are common to the practice of science.
While the larger moose diet study is expected to be published at some point, I don’t expect the DNA sequencing portion of the project to be included in the publication.
Chen, S., H. Yao, J. Han, C. Liu, J. Song, L. Shi, Y. Zhu, X. Ma, T. Gao, X. Pang, K. Luo, Y. Li, X. Li, X. Jia, Y. Lin, and C. Leon. 2010. Validation of the ITS2 Region as a Novel DNA Barcode for Identifying Medicinal Plant Species. PLOS ONE 5:e8613.